SEES:lab awarded grant by the James S. McDonnell Foundation
The SEES:lab has been granted a Research Award from the James S. McDonnell Foundation. The award, for an amount of $445,663, is one of 11 in the Program Area Studying Complex Systems.
Summary of the proposal
Recent technological advances have boosted our ability to gather information on a variety of complex processes occurring at scales from the cellular to the planetary. At a cellular scale, the number of unicellular organisms whose genome has been completely sequenced has grown exponentially in the last few years. For each of these organisms, we know with remarkable accuracy the exact sequence of nucleotides in their DNA-in other words, we know their bio- chemical "blueprint." Unfortunately, our understanding of cellular processes has not grown proportionally. For example, experimental evidence demonstrates that only 20% of the genes in yeast are essential for survival, but the reasons why these genes are essential while others are not are not generally known.
Similarly, at the scale of global socio-economic systems, despite a fast increase in computational capabilities to process global-scale information, our predictive power has not increased at the same pace. For example, nowadays it is possible to track financial and commercial relationships between companies, but when a global crisis develops little is known about what will be the ultimate consequences or about which countries will be most affected.
One of the reasons behind the disparity between the amount information available to us and our understanding of these problems is that in complex systems, such as cells and economies, individual components interact with each other through complex networks of interactions that are neither totally regular nor totally random. The reason why real-world networks are non- trivial is that any bias, however small, in the way components establish connections gives rise to structural correlations. This makes understanding complex systems challenging but, at the same time, it means that each network contains, hidden within its structure, important cues about how the system operates and evolves.
The goal of our research is to develop a theoretical framework and computational tools that will help us, for example, understand a certain cellular function from limited knowledge about the interactions between the proteins involved; or uncover novel gene-disease associations from the relatively few associations we already know; or determine which interactions in an economy are the most critical to avoid systemic crisis.
Although in the last ten years significant progress has been made in the study of networks, we are still far from the ultimate goals of: (i) fully characterizing real-world complex networks; (ii) understanding the precise mechanisms responsible for the observed topologies; and (iii) evaluating the impact of the structure of the network on the dynamics of the system.
Consider the case of the human proteome-the network formed by all proteins expressed in human cells and all the interactions between these proteins. The proteome holds the promise of uncovering the fundamental processes occurring within normal and diseased cells. Ultimately, there is reason to trust that fully characterizing the proteome will enable us to design better diagnostic tools, to identify more suitable drug targets (more effective and with less side effects), and to design better therapies-in short, to significantly improve our quality of life. These prospects may justify the billion-dollar estimated cost of the human proteome project.
The human proteome project, however, also provides an illustration of the challenges ahead of us. Proteome-charting relies on high-throughput techniques whose accuracies were estimated in 2002 to be below 20%. Even after spending millions of dollars and using such "quick-and- dirty" methods, current estimates suggest that a dismal 0.3% of all the physical protein-protein interactions in human proteome have been uncovered so far. The first challenge is, therefore, to obtain reliable reconstructions of this network.
To grasp the magnitude of the second challenge, imagine that we somehow manage to obtain the remaining 99.7% of the human proteome in the near future-we are then left with about 25,000 proteins and 650,000 interactions between them. How are we supposed to identify drug targets from such a mass of data? Which are the basic organizational principles of the network? Which are the key players (proteins and interactions)? Where do we even start looking?
These two challenges (data reliability and information overload) are not specific to the human proteome or even to networks in systems biology. Consider the problem of charting how species interact in ecosystems and how this impacts the propagation of contaminants and other types of ecological perturbations; or, how people commute and travel every day and the impact this has on the spread of epidemics; or the role of the network of inter-organization financial relationships and its role in the making and propagation of a global economic crisis.
As daunting as the challenges may seem, we are now closer to their solution thanks to a decade of network research, and especially to recent progress in understanding the group structure of complex networks. Indeed, individual components of complex systems can be classified into groups, and group membership determines the pattern of interactions of each component. As an illustrative example, consider the air transportation network, in which nodes represent cities and connections represent direct flights between cities-two cities in the same continent are more likely to be connected to one other than two cities in different continents.
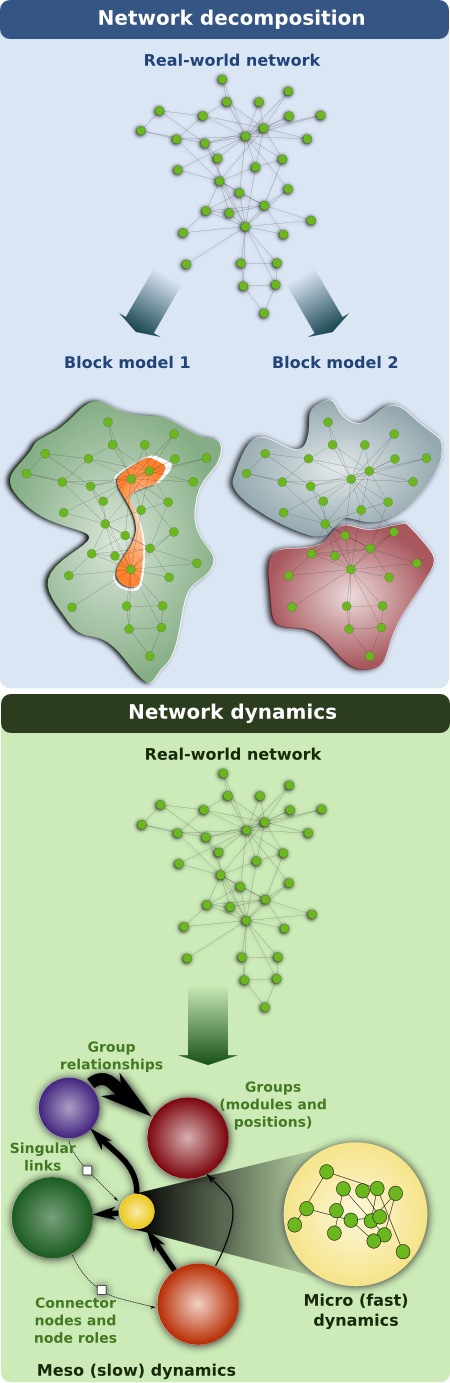
An important consideration is that, in general, there are multiple relevant groupings of the components of the system. For example, in the air transportation network cities can also be grouped according to their role, so that international hubs like Paris or New York are likely to be connected to each other, regardless of whether they are in different continents. The "geographic" and the "role" descriptions of the air transportation network convey different connection mechanisms that reflect the historical and economic pressures that have shaped the network: the network we observe is the result of the superposition of these two mechanisms (and probably others).
Unlike for the air transportation network, we do not usually know a priori which are the relevant groupings of the components of a system (say, proteins in the human proteome). Importantly, we have recently shown that using statistical inference one can identify and sample among groupings with significant explanatory power. This approach opens the door to tackle the two network challenges described above. First, by identifying the most relevant groupings we can point out connections that deviate from the leading group-to-group patterns, which may correspond to observation errors or to connections with special functional requirements (and with special dynamical roles). Second, we can use the most relevant groupings to make educated guesses as to what connections are missing from our reconstructions and, ultimately, to reconstruct whole networks from partial and sparse observations. Finally, the most relevant groupings provide optimal coarse-grained descriptors of the networks, thus helping us solve the "information overload" challenge.
After a decade of network research, the time has come to provide answers that will usher in real breakthroughs in our understanding of complex systems. These breakthroughs will not come from a single approach; ours builds upon some of the knowledge accumulated during these years, and holds the promise to provide a connection between topology and dynamics using a single comprehensive framework.